# Create sample data for Bland-Altman plot (paired measurements)
set.seed(123)
n <- 50
method1 <- rnorm(n, mean = 75, sd = 10)
# Method 2 has slight systematic bias (+2) and more variability
method2 <- method1 + 2 + rnorm(n, mean = 0, sd = 3)
# Calculate Bland-Altman metrics
mean_score <- (method1 + method2) / 2
difference <- method1 - method2
mean_diff <- mean(difference)
sd_diff <- sd(difference)
loa_upper <- mean_diff + 1.96 * sd_diff
loa_lower <- mean_diff - 1.96 * sd_diff
# Create Bland-Altman plot
bland_altman_data <- data.frame(mean_score, difference)
ggplot(bland_altman_data, aes(x = mean_score, y = difference)) +
geom_point(color = "black", alpha = 0.7) +
geom_hline(yintercept = mean_diff, color = "gray40", linetype = "solid", linewidth = 1) +
geom_hline(yintercept = loa_upper, color = "gray60", linetype = "dashed", linewidth = 0.8) +
geom_hline(yintercept = loa_lower, color = "gray60", linetype = "dashed", linewidth = 0.8) +
geom_hline(yintercept = 0, color = "black", linetype = "dotted", linewidth = 0.5) +
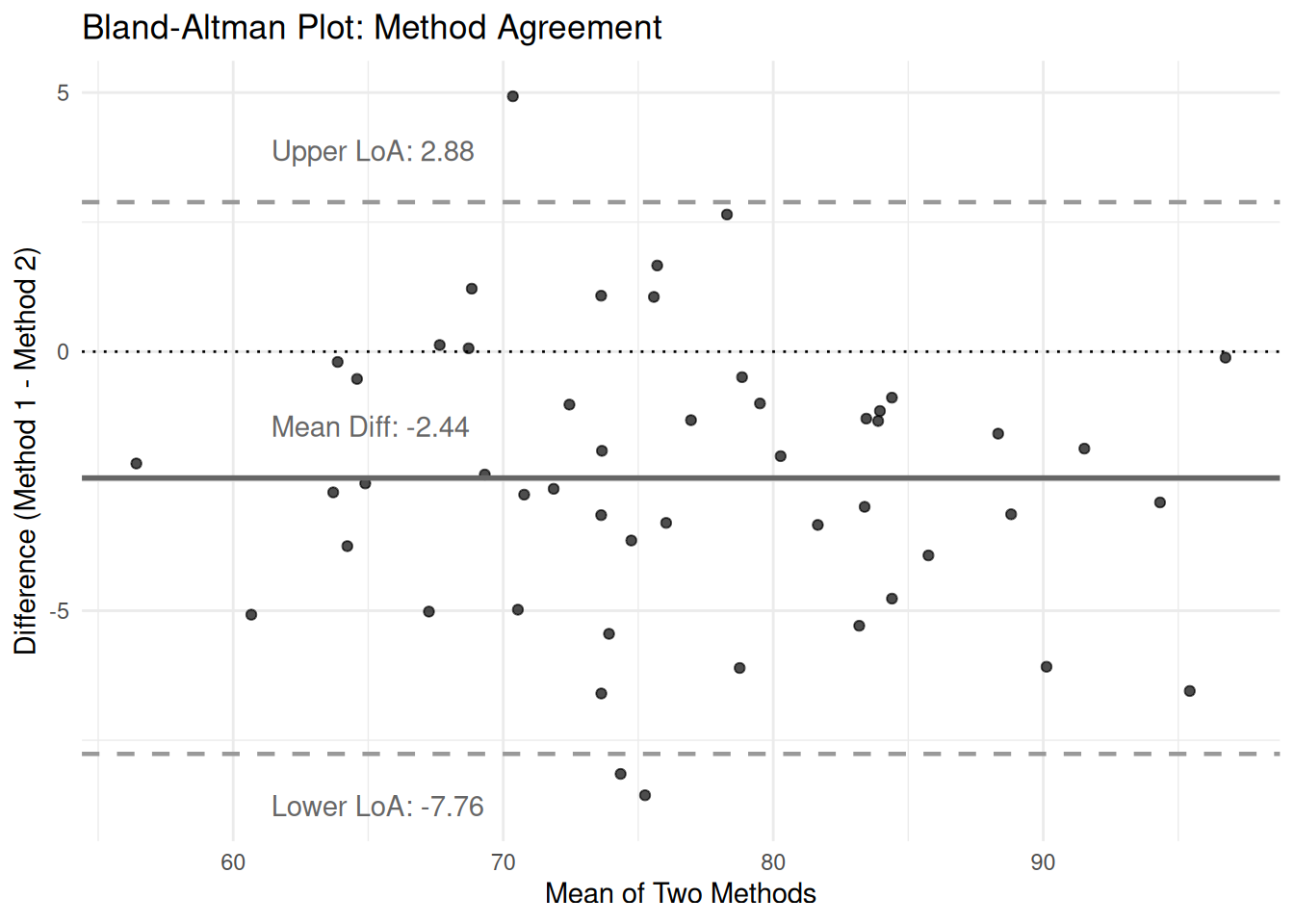
labs(title = "Bland-Altman Plot: Method Agreement",
x = "Mean of Two Methods",
y = "Difference (Method 1 - Method 2)") +
theme_minimal() +
annotate("text", x = min(mean_score) + 5, y = loa_upper + 1,
label = paste("Upper LoA:", round(loa_upper, 2)), hjust = 0, color = "gray40") +
annotate("text", x = min(mean_score) + 5, y = loa_lower - 1,
label = paste("Lower LoA:", round(loa_lower, 2)), hjust = 0, color = "gray40") +
annotate("text", x = min(mean_score) + 5, y = mean_diff + 1,
label = paste("Mean Diff:", round(mean_diff, 2)), hjust = 0, color = "gray40")